- ImageCLEF 2026
- LifeCLEF 2026
- ImageCLEF 2025
- LifeCLEF 2025
- ImageCLEF 2024
- LifeCLEF 2024
- ImageCLEF 2023
- LifeCLEF 2023
- ImageCLEF 2022
- LifeCLEF2022
- ImageCLEF 2021
- LifeCLEF 2021
- ImageCLEF 2020
- LifeCLEF 2020
- ImageCLEF 2019
- LifeCLEF 2019
- ImageCLEF 2018
- LifeCLEF 2018
- ImageCLEF 2017
- LifeCLEF2017
- ImageCLEF 2016
- LifeCLEF 2016
- ImageCLEF 2015
- LifeCLEF 2015
- ImageCLEF 2014
- LifeCLEF 2014
- ImageCLEF 2013
- ImageCLEF 2012
- ImageCLEF 2011
- ImageCLEF 2010
- ImageCLEF 2009
- ImageCLEF 2008
- ImageCLEF 2007
- ImageCLEF 2006
- ImageCLEF 2005
- ImageCLEF 2004
- ImageCLEF 2003
- Publications
- Old resources
You are here
ImageCLEFmed Caption
Welcome to the 3rd edition of the Caption Task!
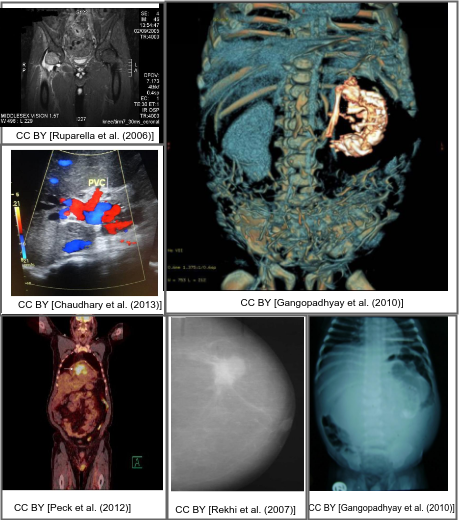
Motivation
Interpreting and summarizing the insights gained from medical images such as radiology output is a time-consuming task that involves highly trained experts and often represents a bottleneck in clinical diagnosis pipelines.
Consequently, there is a considerable need for automatic methods that can approximate this mapping from visual information to condensed textual descriptions. The more image characteristics are known, the more structured are the radiology scans and hence, the more efficient are the radiologists regarding interpretation. We work on the basis of a large-scale collection of figures from open access biomedical journal articles (PubMed Central). All images in the training data are accompanied by UMLS concepts extracted from the original image caption.
Lessons learned:
- In the first and second editions of this task, held at ImageCLEF 2017 and ImageCLEF 2018, participants noted a broad variety of content and situation among training images. For this year, the training data is reduced solely to radiology images
- A large number of concepts was used in the previous years. This year, the captions are first processed before concept extraction, hence leading to a reduced number of concepts
- As uncertainty regarding additional source was noted, we will clearly separate systems using exclusively the official training data from those that incorporate additional sources of evidence
News
- 08.11.2018: website goes live
- 25.11.2018: registration open at crowdAI
- 15.01.2019: development data released at crowdAI
- 18.03.2019: test data released at crowdAI
- 29.04.2019: evaluate-f1.py updated
- 29.04.2019: submission deadline extended to 06.05.2019 (AoE)
- 19.05.2019: working notes instruction added
- 23.05.2019: working notes deadline extended to 27.05.2019 (UTC)
Task Description
Concept Detection Task
The first step to automatic image captioning and scene understanding is identifying the presence and location of relevant concepts in a large corpus of medical images. Based on the visual image content, this subtask provides the building blocks for the scene understanding step by identifying the individual components from which captions are composed. The concepts can be further applied for context-based image and information retrieval purposes.
Evaluation is conducted in terms of set coverage metrics such as precision, recall, and combinations thereof. This task will be run using a subset of the Radiology Objects in COntext (ROCO) dataset [1].
Data
From the PubMed Open Access subset containing 1,828,575 archives, a total number of 6,031,814
image - caption pairs were extracted. To focus on radiology images and non-compound figures, automatic filtering with deep learning systems as well as manual revisions were applied, reducing the dataset to 70,786 radiology images of several medical imaging modalities.
NOTE: If the usage of an additional source for training is intended, it should not be a subset of PubMed Central Open Access (archiving date: 01.02.2018 - 01.02.2019), to avoid an overlap with the test data.
Evaluation Methodology
Evaluation is conducted in terms of F1 scores between system predicted and ground truth concepts, using the following methodology and parameters:
- The default implementation of the Python scikit-learn (v0.17.1-2) F1 scoring method is used. It is documented here.
- A Python (3.x) script loads the candidate run file, as well as the ground truth (GT) file, and processes each candidate-GT concept sets
- For each candidate-GT concept set, the y_pred and y_true arrays are generated. They are binary arrays indicating for each concept contained in both candidate and GT set if it is present (1) or not (0).
- The F1 score is then calculated. The default 'binary' averaging method is used.
- All F1 scores are summed and averaged over the number of elements in the test set (10'000), giving the final score.
The ground truth for the test set was generated based on the UMLS Full Release 2017AB.
NOTE: The source code of the evaluation tool is available here. It must be executed using Python 3.x, on a system where the scikit-learn (>= v0.17.1-2) Python library is installed. The script should be run like this:
/path/to/python3 evaluate-f1.py /path/to/candidate/file /path/to/ground-truth/file
Preliminary Schedule
- 05.11.2018: Registration opens for all ImageCLEF tasks (until 26.04.2019)
- 15.01.2019: Development data release starts
- 18.03.2019: Test data release starts
01.05.201906.05.2019: Deadline for submitting the participants runs- 13.05.2019: Release of the processed results by the task organizers
24.05.201927.05.2019: Deadline for submission of working notes papers by the participants- 07.06.2019: Notification of acceptance of the working notes papers
- 28.06.2019: Camera-ready working notes papers
- 09-12.09.2019: CLEF 2019, Lugano, Switzerland
Participant Registration
Please refer to the general ImageCLEF registration instructions
Submission Instructions
Please note that each group is allowed a maximum of 10 runs per subtask.
For the submission of the concept detection task we expect the following format:
- <Figure-ID><TAB><Concept-ID-1>;<Concept-ID-2>;<Concept-ID-n>
e.g.:
- ROCO_CLEF_41341 C0033785;C0035561
- ROCO_CLEF_07563 C0043299;C1306645;C1548003;C1962945
You need to respect the following constraints:
- The separator between the figure ID and the concepts has to be a tabular whitespace
- The separator between the UMLS concepts has to be a semicolon (;)
- Each figure ID of the test set must be included in the submitted file exactly once (even if there are not concepts)
- The same concept cannot be specified more than once for a given figure ID
- The maximum number of concepts per image is 100
Results
| Group Name | Submission Run | F1 Score | Rank |
|---|---|---|---|
| AUEB NLP Group | s2_results.csv | 0.2823094 | 1 |
| AUEB NLP Group | ensemble_avg.csv | 0.2792511 | 2 |
| AUEB NLP Group | s1_results.csv | 0.2740204 | 3 |
| damo | test_cat_xi.txt | 0.2655099 | 4 |
| AUEB NLP Group | s3_results.csv | 0.2639952 | 5 |
| damo | test_results.txt | 0.2613895 | 6 |
| damo | first_concepts_detection_result_check.txt | 0.2316484 | 7 |
| ImageSem | F1TOP1.txt | 0.2235690 | 8 |
| ImageSem | F1TOP2.txt | 0.2227917 | 9 |
| ImageSem | F1TOP5_Pmax.txt | 0.2216225 | 10 |
| ImageSem | F1TOP3.txt | 0.2190201 | 11 |
| ImageSem | 07Comb_F1Top1.txt | 0.2187337 | 12 |
| ImageSem | F1TOP5_Rmax.txt | 0.2147437 | 13 |
| damo | test_tran_all.txt | 0.2134523 | 14 |
| damo | test_cat.txt | 0.2116252 | 15 |
| UA.PT_Bioinformatics | simplenet.csv | 0.2058640 | 16 |
| richard_ycli | testing_result.txt | 0.1952310 | 17 |
| ImageSem | 08Comb_Pmax.txt | 0.1912173 | 18 |
| UA.PT_Bioinformatics | simplenet128x128.csv | 0.1893430 | 19 |
| UA.PT_Bioinformatics | mix-1100-o0-2019-05-06_1311.csv | 0.1825418 | 20 |
| UA.PT_Bioinformatics | aae-1100-o0-2019-05-02_1509.csv | 0.1760092 | 21 |
| Sam Maksoud | TRIAL_1.txt | 0.1749349 | 22 |
| richard_ycli | testing_result.txt | 0.1737527 | 23 |
| UA.PT_Bioinformatics | ae-1100-o0-2019-05-02_1453.csv | 0.1715210 | 24 |
| UA.PT_Bioinformatics | cedd-1100-o0-2019-05-03_0937-trim.csv | 0.1667884 | 25 |
| AI600 | ai600_result_weighing_1557061479.txt | 0.1656261 | 26 |
| Sam Maksoud | TRIAL_18.txt | 0.1640647 | 27 |
| richard_ycli | testing_result_run4.txt | 0.1633958 | 28 |
| AI600 | ai600_result_weighing_1557059794.txt | 0.1628424 | 29 |
| richard_ycli | testing_result_run3.txt | 0.1605645 | 30 |
| AI600 | ai600_result_weighing_1557107054.txt | 0.1603341 | 31 |
| AI600 | ai600_result_weighing_1557062212.txt | 0.1588862 | 32 |
| AI600 | ai600_result_weighing_1557062494.txt | 0.1562828 | 33 |
| AI600 | ai600_result_weighing_1557107838.txt | 0.1511505 | 34 |
| richard_ycli | testing_result_run2.txt | 0.1467212 | 35 |
| MacUni-CSIRO | run1FinalOutput.txt | 0.1435435 | 36 |
| AI600 | ai600_result_rgb_1556989393.txt | 0.1345022 | 37 |
| UA.PT_Bioinformatics | simplenet64x64.csv | 0.1279909 | 38 |
| UA.PT_Bioinformatics | resnet19-cnn.csv | 0.1269521 | 39 |
| ImageSem | 09Comb_Rmax_new.txt | 0.1121941 | 40 |
| damo | test_att_3_rl_best.txt | 0.0590448 | 41 |
| damo | test_rl_5_result_check.txt | 0.0584684 | 42 |
| damo | test_tran_rl_5.txt | 0.0567311 | 43 |
| damo | test_tran_10.txt | 0.0536554 | 44 |
| pri2si17 | submission_1.csv | 0.0496821 | 45 |
| AILAB | results_v3.txt | 0.0202243 | 46 |
| AILAB | results_v1.txt | 0.0198960 | 47 |
| AILAB | results_v2.txt | 0.0162458 | 48 |
| pri2si17 | submission_3.csv | 0.0141422 | 49 |
| AILAB | results_v4.txt | 0.0126845 | 50 |
| LIST | denseNet_pred_all_0.55.txt | 0.0013269 | 51 |
| ImageSem | yu_1000_inception_v3_top6.csv | 0.0009450 | 52 |
| ImageSem | yu_1000_resnet_152_top6.csv | 0.0008925 | 53 |
| LIST | denseNet_pred_all_0.6.txt | 0.0003665 | 54 |
| LIST | denseNet_pred_all.txt | 0.0003400 | 55 |
| LIST | predictionBR(LR).txt | 0.0002705 | 56 |
| LIST | denseNet_pred_all_0.6_50_0.04(max if null).txt | 0.0002514 | 57 |
| LIST | predictionCC(LR).txt | 0.0002494 | 58 |
| AILAB | results_v0.txt | 0 | - |
| pri2si17 | submission_2.csv | 0 | - |
CEUR Working Notes
- All participating teams with atleast one graded submission, regardless of F1 score, should submit a CEUR working notes paper.
- The working notes paper should be submitted using this link:
https://easychair.org/conferences/?conf=clef2019
Click on "enter as an author", then select track "ImageCLEF - Multimedia Retrieval in CLEF".
Add author information, paper title/abstract, keywords, select "Task 3 - ImageCLEFmedical" and upload your working notes paper as pdf.
Citations
When referring to the ImageCLEFmed 2019 concept detection task general goals, general results, etc. please cite the following publication which will be published by September 2019:
- Obioma Pelka, Christoph M. Friedrich, Alba García Seco de Herrera and Henning Müller. Overview of the ImageCLEFmed 2019 Concept Detection Task, CEUR Workshop Proceedings (CEUR- WS.org), ISSN 1613-0073, http://ceur-ws.org/Vol-2380/.
- BibTex:
@Inproceedings{ImageCLEFmedConceptOverview2019,
author = {Pelka, Obioma and Friedrich, Christoph M and Garc\'ia Seco de Herrera, Alba and M\"uller, Henning},
title = {Overview of the {ImageCLEFmed} 2019 Concept Prediction Task},
booktitle = {CLEF2019 Working Notes},
series = {{CEUR} Workshop Proceedings},
year = {2019},
volume = {2380},
publisher = {CEUR-WS.org $<$http://ceur-ws.org$>$},
pages = {},
month = {September 09-12},
address = {Lugano, Switzerland},
}
When referring to the ImageCLEF 2019 task in general, please cite the following publication to be published by September 2019:
- BibTex:
@InProceedings{ImageCLEF19,
author = {Bogdan Ionescu and Henning M\"uller and Renaud P\'{e}teri
and Yashin Dicente Cid and Vitali Liauchuk and Vassili Kovalev and
Dzmitri Klimuk and Aleh Tarasau and Asma Ben Abacha and Sadid A. Hasan
and Vivek Datla and Joey Liu and Dina Demner-Fushman and Duc-Tien
Dang-Nguyen and Luca Piras and Michael Riegler and Minh-Triet Tran and
Mathias Lux and Cathal Gurrin and Obioma Pelka and Christoph M.
Friedrich and Alba Garc\'ia Seco de Herrera and Narciso Garcia and
Ergina Kavallieratou and Carlos Roberto del Blanco and Carlos Cuevas
Rodr\'{i}guez and Nikos Vasillopoulos and Konstantinos Karampidis and
Jon Chamberlain and Adrian Clark and Antonio Campello},
title={{ImageCLEF 2019}: Multimedia Retrieval in Medicine, Lifelogging, Security and Nature},
booktitle={Experimental IR Meets Multilinguality, Multimodality, and Interaction},
series = {Proceedings of the Tenth International Conference of the CLEF Association (CLEF 2019)},
year = {2019},
volume = {},
publisher = {{LNCS} Lecture Notes in Computer Science, Springer},
pages = {},
month = {September 09-12},
address = {Lugano, Switzerland},}
Contact
- Obioma Pelka <obioma.pelka(at)fh-dortmund.de>, University of Applied Sciences and Arts Dortmund, Germany
- Christoph M. Friedrich <christoph.friedrich(at)fh-dortmund.de>, University of Applied Sciences and Arts Dortmund, Germany
- Alba García Seco de Herrera <alba.garcia(at)essex.ac.uk>,University of Essex, UK
- Henning Müller <henning.mueller(at)hevs.ch>, University of Applied Sciences Western Switzerland, Sierre, Switzerland
Join our mailing list: https://groups.google.com/d/forum/imageclefcaption
Follow @imageclef
Acknowledgments
[1] O. Pelka, S. Koitka, J. Rückert, F. Nensa und C. M. Friedrich „Radiology Objects in COntext (ROCO): A Multimodal Image Dataset“, Proceedings of the MICCAI Workshop on Large-scale Annotation of Biomedical data and Expert Label Synthesis (MICCAI LABELS 2018), Granada, Spain, September 16, 2018, Lecture Notes in Computer Science (LNCS) Volume 11043, Page 180-189, DOI: 10.1007/978-3-030-01364-6_20, Springer Verlag, 2018.
| Attachment | Size |
|---|---|
| 608.47 KB |

