- ImageCLEF 2025
- LifeCLEF 2025
- ImageCLEF 2024
- LifeCLEF 2024
- ImageCLEF 2023
- LifeCLEF 2023
- ImageCLEF 2022
- LifeCLEF2022
- ImageCLEF 2021
- LifeCLEF 2021
- ImageCLEF 2020
- LifeCLEF 2020
- ImageCLEF 2019
- LifeCLEF 2019
- ImageCLEF 2018
- LifeCLEF 2018
- ImageCLEF 2017
- LifeCLEF2017
- ImageCLEF 2016
- LifeCLEF 2016
- ImageCLEF 2015
- LifeCLEF 2015
- ImageCLEF 2014
- LifeCLEF 2014
- ImageCLEF 2013
- ImageCLEF 2012
- ImageCLEF 2011
- ImageCLEF 2010
- ImageCLEF 2009
- ImageCLEF 2008
- ImageCLEF 2007
- ImageCLEF 2006
- ImageCLEF 2005
- ImageCLEF 2004
- ImageCLEF 2003
- Publications
- Old resources
You are here
PlantCLEF 2021

| News |
| A direct link to the overview of the task: Overview of PlantCLEF 2021: cross-domain plant identification, Hervé Goëau, Pierre Bonnet, Alexis Joly, LifeCLEF 2021 working notes, Bucharest, Romania |
| Link to the data (same data as 2020 apart from traits data PlantCLEF 2021 Traits data PlantCLEF 2020-21 Training dataset (description) PlantCLEF 2020-21 Training dataset (link) PlantCLEF2020-21 test dataset and training dataset (alternative to zenodo) |
Motivation
For several centuries, botanists have collected, catalogued and systematically stored plant specimens in herbaria. These physical specimens are used to study the variability of species, their phylogenetic relationship, their evolution, or phenological trends. One of the key step in the workflow of botanists and taxonomists is to find the herbarium sheets that correspond to a new specimen observed in the field. This task requires a high level of expertise and can be very tedious. Developing automated tools to facilitate this work is thus of crucial importance. More generally, this will help to convert these invaluable centuries-old materials into FAIR data.
Data collection
The task will rely on a large collection of more than 60,000 herbarium sheets that were collected in French Guyana (i.e. from the Herbier IRD de Guyane ) and digitized in the context of the e-ReColNat project. iDigBio (the US National Resource for Advancing Digitization of Biodiversity Collections) hosts millions of images of herbarium specimens. Several tens of thousands of these images, illustrating the French Guyana flora, will be used for the PlantCLEF task this year. A valuable asset of this collection is that several herbarium sheets are accompanied by a few pictures of the same specimen in the field. For the test set, we will use in-the-field pictures coming different sources including Pl@ntNet and Encyclopedia of Life.
The training dataset of the LifeCLEF2021 Plant Identification challenge will be based on the same visual data used during the previous LifeCLEF 2020 Plant Identification challenge but will also introduce new data related to 5 "traits" covering exhaustively all the 1000 species of the challenge. Traits are a very valuable information that can potentially help improve prediction models. Indeed, it can be assumed that species which share the same traits also share to some extent common visual appearances. This information can then potentially be used to guide the learning of a model through auxiliary loss functions for instance.
These data were collected through the Encyclopedia of Life API. The 5 most exhaustive traits ("plant growth form", "habitat", "plant lifeform", "trophic guild" and "woodiness") were verified and completed by experts of the Guyanese flora, so that each of the 1000 species have a value for each trait.
Task description
The challenge will be evaluated as a cross-domain classification task. The training set will consist of herbarium sheets whereas the test set will be composed of field pictures. To enable learning a mapping between the herbarium sheets domain and the field pictures domain, we will provide both herbarium sheets and field pictures for a subset of species. The metrics used for the evaluation of the task will be the classification accuracy and the Mean Reciprocal Rank.
How to participate ?
Follow this link to the AIcrowd platform LifeCLEF 2021 Plant
- Each participant has to register on AIcrowd (https://www.aicrowd.com/) with username, email and password. A representative team name should be used
as username. -
In order to be compliant with the CLEF requirements, participants also have to fill in the following additional fields on their profile:
- First name
- Last name
- Affiliation
- Address
- City
- Country
-
This information will not be publicly visible and will be exclusively used to contact you and to send the registration data to CLEF, which is the main organizer of all CLEF labs. Once set up, participants will have access to the dataset tab on the challenge's page. A LifeCLEF participant will be considered as registered for a task as soon as he/she has downloaded a file of the task's dataset via the dataset tab of the challenge.
Reward
The winner of each of the four LifeCLEF 2020 challenges will be offered a cloud credit grants of 5k USD as part of Microsoft's AI for earth program.

Results
A total of 4 participating groups submitted 29 runs. Thanks to all of you for your efforts!
| Team run name | Aicrowd name | Filename | MRR (whole test set) | MRR (sub-set of the test set related to species with few training photos in the field) |
|---|---|---|---|---|
| Neuon AI Run 7 | holmes_chang | submissionformat_11_run8_run1_run11_run13_run14.csv | 0.181 | 0.158 |
| Neuon AI Run 10 | holmes_chang | submissionformat_14_run14t_run1_run11_run13_run14.csv | 0.176 | 0.153 |
| Neuon AI Run 8 | holmes_chang | submissionformat_12_run8_run1_run11.csv | 0.169 | 0.15 |
| Neuon AI Run 2 | holmes_chang | 2_mixed_run13_run14_precrop_mean_emb_cosine_similarity_inverse_weight_flip_crop.csv | 0.152 | 0.117 |
| Neuon AI Run 9 | holmes_chang | submissionformat_13_run8_run1_run11_run13_run14_e.csv | 0.147 | 0.129 |
| Neuon AI Run 6 | holmes_chang | submissionformat_10_run8_run13_run14.csv | 0.143 | 0.126 |
| Neuon AI Run 5 | holmes_chang | submissionformat_9_run1_run13_run14.csv | 0.137 | 0.116 |
| Neuon AI Run 4 | holmes_chang | submissionformat_5_run6_run13_run14.csv | 0.088 | 0.073 |
| Neuon AI Run 1 | holmes_chang | run1_cont_precrop_mean_emb_cosine_similarity_inverse_weight_flip_crop.csv | 0.071 | 0.066 |
| LU Run 9 | heaven | Submission_2021_3.56_3.35_n_times.txt | 0.065 | 0.037 |
| Domain Run 8 | Domain_run | Submission_2021_3.7.txt | 0.065 | 0.037 |
| LU Run 8 | heaven | Submission_2021_3.56_raw.txt | 0.065 | 0.037 |
| LU Run 7 | heaven | Submission_2021_3.56_3.35.txt | 0.063 | 0.04 |
| Domain Run 4 | Domain_run | Submission_2021_3.txt | 0.063 | 0.046 |
| Neuon AI Run 3 | holmes_chang | submissionformat_3_run6.csv | 0.06 | 0.056 |
| Domain Run 6 | Domain_run | Submission_2021_3.35.txt | 0.06 | 0.039 |
| LU Run 6 | heaven | predictions_2021_3.5921463632307007_weight.txt | 0.057 | 0.042 |
| To Be Run 2 | To_be | predictions_2021_3.525211958946899.txt | 0.056 | 0.038 |
| LU Run 4 | heaven | predictions_2021_52.343750.095351815_weight.txt | 0.051 | 0.031 |
| LU Run 3 | heaven | predictions_2021_46.8750.09727478_weight.txt | 0.05 | 0.026 |
| To Be Run 1 | To_be | predictions_20021_49.218750.09158182.txt | 0.045 | 0.022 |
| Domain Run 5 | Domain_run | Submission_2021_408.txt | 0.038 | 0.011 |
| LU Run 5 | heaven | predictions_20021_50.781250.08264196.txt | 0.037 | 0.022 |
| LU Run 1 | heaven | Submission_2021_4.71_ass.txt | 0.036 | 0.009 |
| LU Run 2 | heaven | Submission_2021_46.8750_all.txt | 0.034 | 0.013 |
| Domain Run 3 | Domain_run | Submission_2.txt | 0.031 | 0.015 |
| Domain Run 1 | Domain_run | Submission_2021_1.txt | 0.019 | 0.009 |
| Domain Run 2 | Domain_run | Submission_2021_2.txt | 0.018 | 0.009 |
| Domain Run 7 | Domain_run | Submission_2021_5.56.txt | 0.006 | 0.003 |
| Organizer's submission 7 | herve.goeau | fsda_classifier_extra_stage_5_final_genus_family_climber_herb_plant_lifeform_shrub_tree_woodiness_best_acc_on_photo_val_7_aug.csv | 0.198 | 0.093 |
| Organizer's submission 6 | herve.goeau | fsda_classifier_extra_stage_5_final_genus_family_climber_herb_shrub_tree_best_acc_on_photo_val_7_aug.csv | 0.192 | 0.091 |
| Organizer's submission 4 | herve.goeau | fsda_classifier_extra_stage_5_final_genus_family_plant_lifeform_best_acc_on_photo_val_7_aug.csv | 0.188 | 0.073 |
| Organizer's submission 5 | herve.goeau | fsda_classifier_extra_stage_5_final_genus_family_woodiness_best_acc_on_photo_val_7_aug.csv | 0.184 | 0.077 |
| Organizer's submission 3 | herve.goeau | fsda_classifier_extra_stage_5_final_genus_family_best_acc_on_photo_val_7_aug.csv | 0.183 | 0.073 |
| Organizer's submission 2 | herve.goeau | fsda_extra_final_single_loop_ep30_aug_7.csv | 0.153 | 0.056 |
| Organizer's submission 1 | herve.goeau | fsda_final_aug_7.csv | 0.052 | 0.042 |
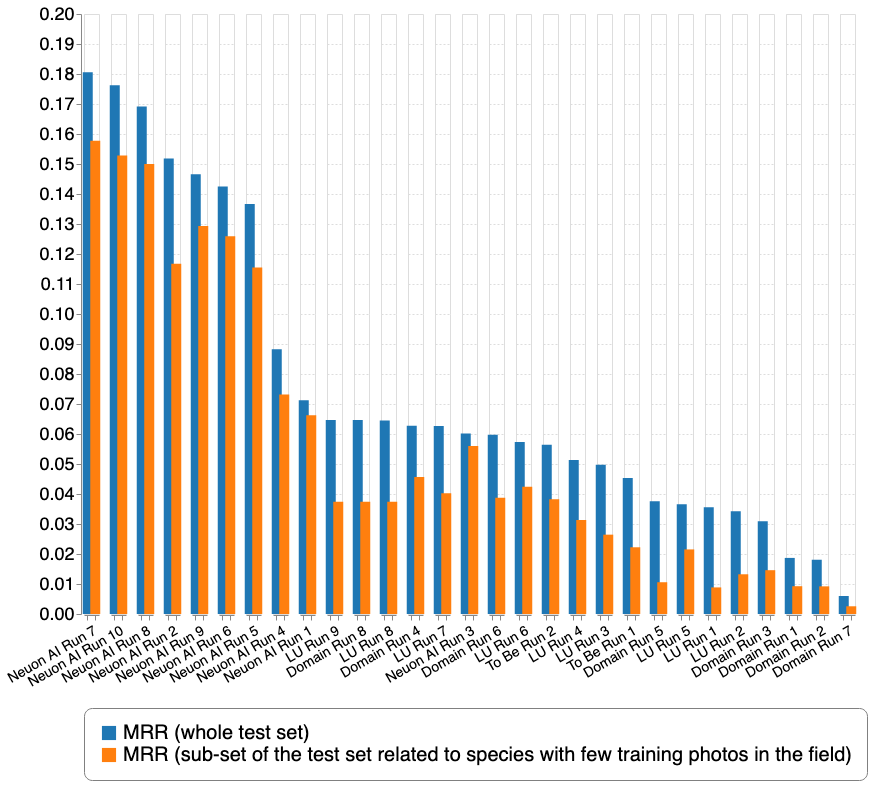
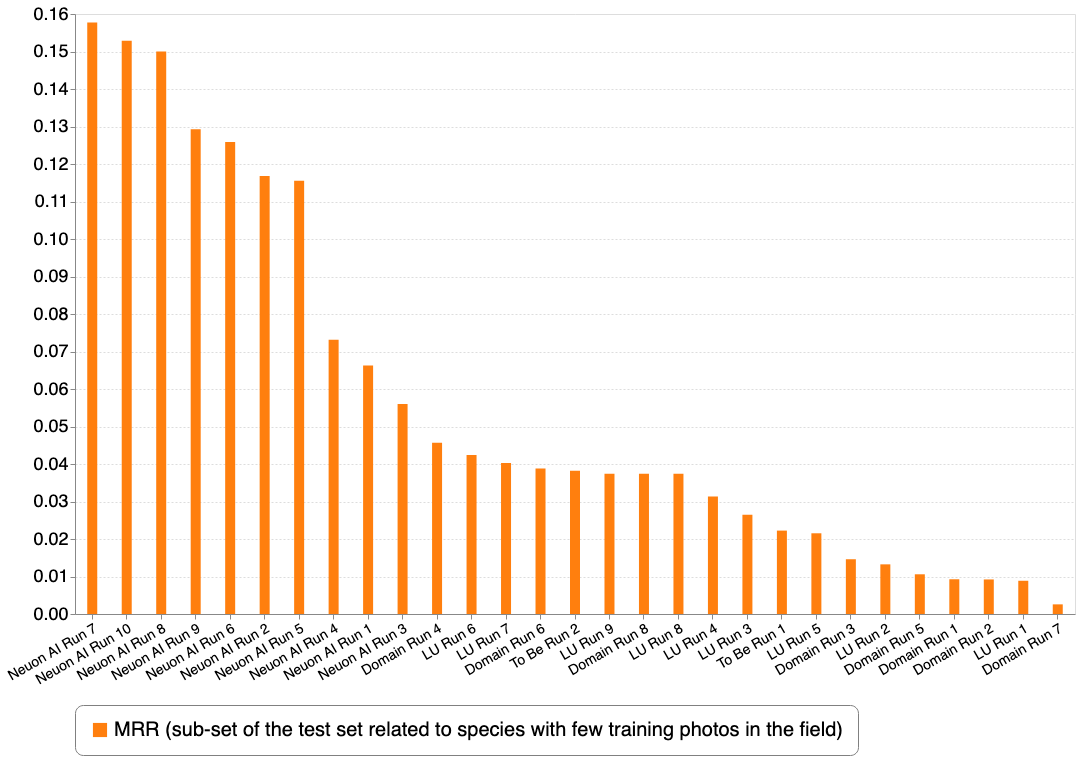
This second graph focuses on the very difficult sub-part of the test set and reorders the submissions according to the second metric.
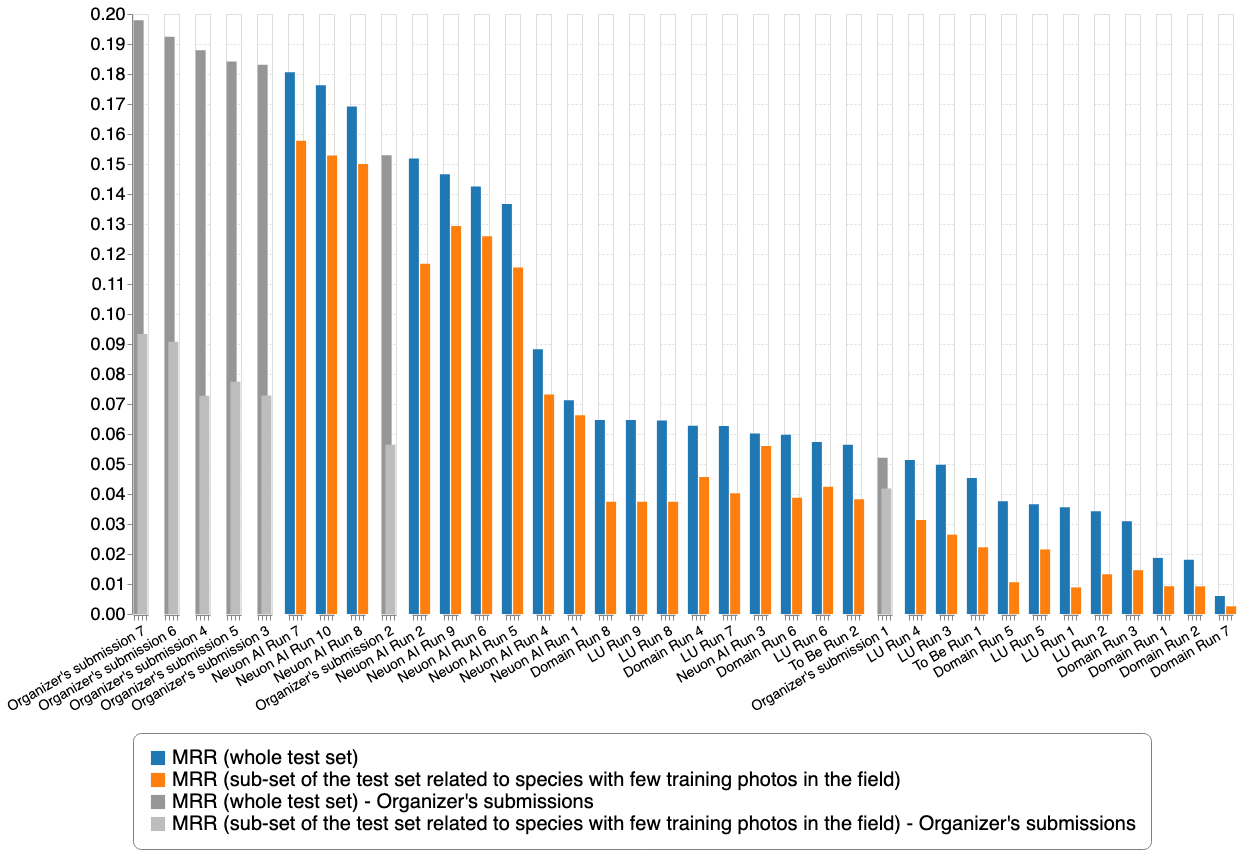
This last graph reports the results with some baselines provided by the organizers in collaboration with Juan Villacis who was ranked first in last year's edition (ITCR PlantNet - aabab).
Credits
| Attachment | Size |
|---|---|
| 71.35 KB | |
| 60.6 KB | |
| 48.7 KB | |
| 179.19 KB | |
| 53.93 KB | |
| 76.88 KB | |
| 68.05 KB | |
| 192.21 KB |
