- ImageCLEF 2025
- LifeCLEF 2025
- ImageCLEF 2024
- LifeCLEF 2024
- ImageCLEF 2023
- LifeCLEF 2023
- ImageCLEF 2022
- LifeCLEF2022
- ImageCLEF 2021
- LifeCLEF 2021
- ImageCLEF 2020
- LifeCLEF 2020
- ImageCLEF 2019
- LifeCLEF 2019
- ImageCLEF 2018
- LifeCLEF 2018
- ImageCLEF 2017
- LifeCLEF2017
- ImageCLEF 2016
- LifeCLEF 2016
- ImageCLEF 2015
- LifeCLEF 2015
- ImageCLEF 2014
- LifeCLEF 2014
- ImageCLEF 2013
- ImageCLEF 2012
- ImageCLEF 2011
- ImageCLEF 2010
- ImageCLEF 2009
- ImageCLEF 2008
- ImageCLEF 2007
- ImageCLEF 2006
- ImageCLEF 2005
- ImageCLEF 2004
- ImageCLEF 2003
- Publications
- Old resources
You are here
GeoLifeCLEF 2024

Schedule
- December 2023: Registration opens for all LifeCLEF challenges Registration is free of charge
- 28 February 2024: Competition Start
- 24 May 2024: Competition Deadline
- 31 May 2024: Deadline for submission of working note papers by participants [CEUR-WS proceedings]
- 21 June 2024: Notification of acceptance of working note papers [CEUR-WS proceedings]
- 8 July 2024: Camera-ready deadline for working note papers.
- 9-12 Sept 2024: CLEF 2024 Grenoble - France
All deadlines are at 11:59 PM CET of the corresponding day unless otherwise stated.
The competition organizers reserve the right to update the contest timeline if they deem it necessary.
Results
All teams that submitted a solution file on a Kaggle and selected runs for private evaluation and their scores are available on Kaggle.
Official competition results are listed on the private and public leaderboards, respectively.
- The 1st place team: webmaking (Anthony Miyaguchi)
- The 2nd place team: AI2Lab(Yichia Chen, Tai Peng, Weihua Li and Chu-Song Chen)
- The 3rd place team: Miss Qiu (Haixu Liu, Zerui Tao, Penghao Jiang, Qiuzhuang Sun and Muyan Wan)
Motivation
Predicting plant species composition and its change in space and time at a fine resolution is useful for many biodiversity management and conservation scenarios, improving species identification and inventory tools, and educational purposes.
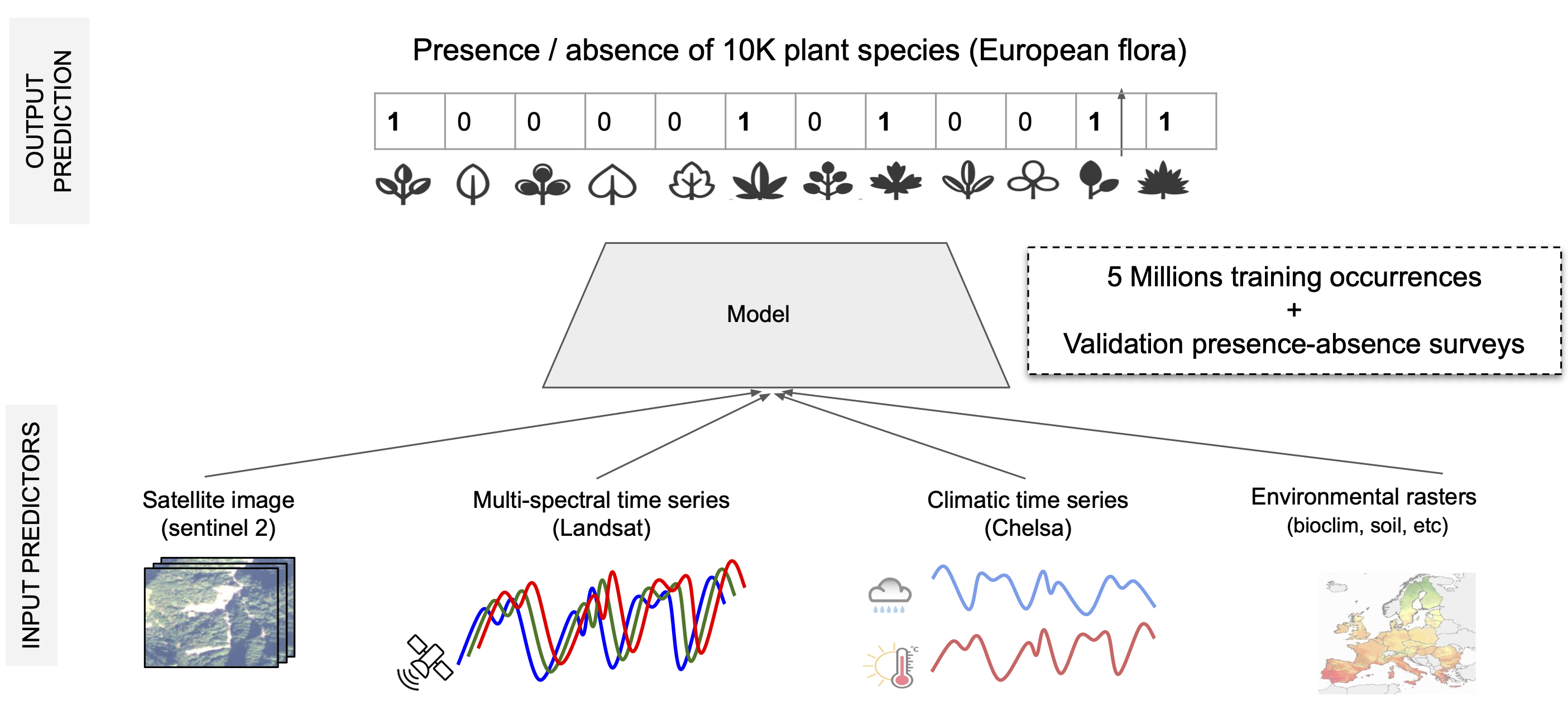
This challenge aims to predict plant species in a given location and time using various possible predictors: satellite images and time series, climatic time series, and other rasterized environmental data: land cover, human footprint, bioclimatic, and soil variables.
To do so, we provide a large-scale training set of about 5M plant occurrences in Europe (single-label, presence-only data) as well as a validation set of about 5K plots and a test set with 20K plots, with all the present species (multi-label, presence-absence data).
The difficulties of the challenge include multi-label learning from single positive labels, strong class imbalance, multi-modal learning, and large-scale.
How to participate?
1. Subscribe to CLEF (GeoLifeCLEF task) by filling this form
2. Go to the Kaggle's GeoLifeCLEF 2024 challenge page and compete: Kaggle challenge
Data collection
The training data comprises species observations and environmental data. Below, we explain the data in detail.
Observations data
The species related training data comprises:
- Presence-Absence (PA) surveys: including around 90 thousand surveys with roughly 10,000 species of the European flora. The presence-absence data (PA) is provided to compensate for the problem of false-absences of PO data and calibrate models to avoid associated biases.
- Presence-Only (PO) occurrences: combines around five million observations from numerous datasets gathered from the Global Biodiversity Information Facility (GBIF, www.gbif.org). This data constitutes the larger piece of the training data and covers all countries of our study area, but it has been sampled opportunistically (without standardized sampling protocol), leading to various sampling biases. The local absence of a species among PO data doesn't mean it is truly absent. An observer might not have reported it because it was difficult to "see" it at this time of the year, to identify it as not a monitoring target, or just unattractive.
There are two CSVs with species occurrence data on the Seafile available for training. The detailed description is provided again on SeaFile in separate ReadME files in relevant folders.
- The PO metadata are available in PresenceOnlyOccurences/GLC24_PO_metadata_train.csv.
- The PA metadata are available in PresenceAbsenceSurveys/GLC24_PA_metadata_train.csv.
Environmental data
Besides species data, we provide spatialized geographic and environmental data as additional input variables (see Figure 1). More precisely, For each species observation location, we provide:
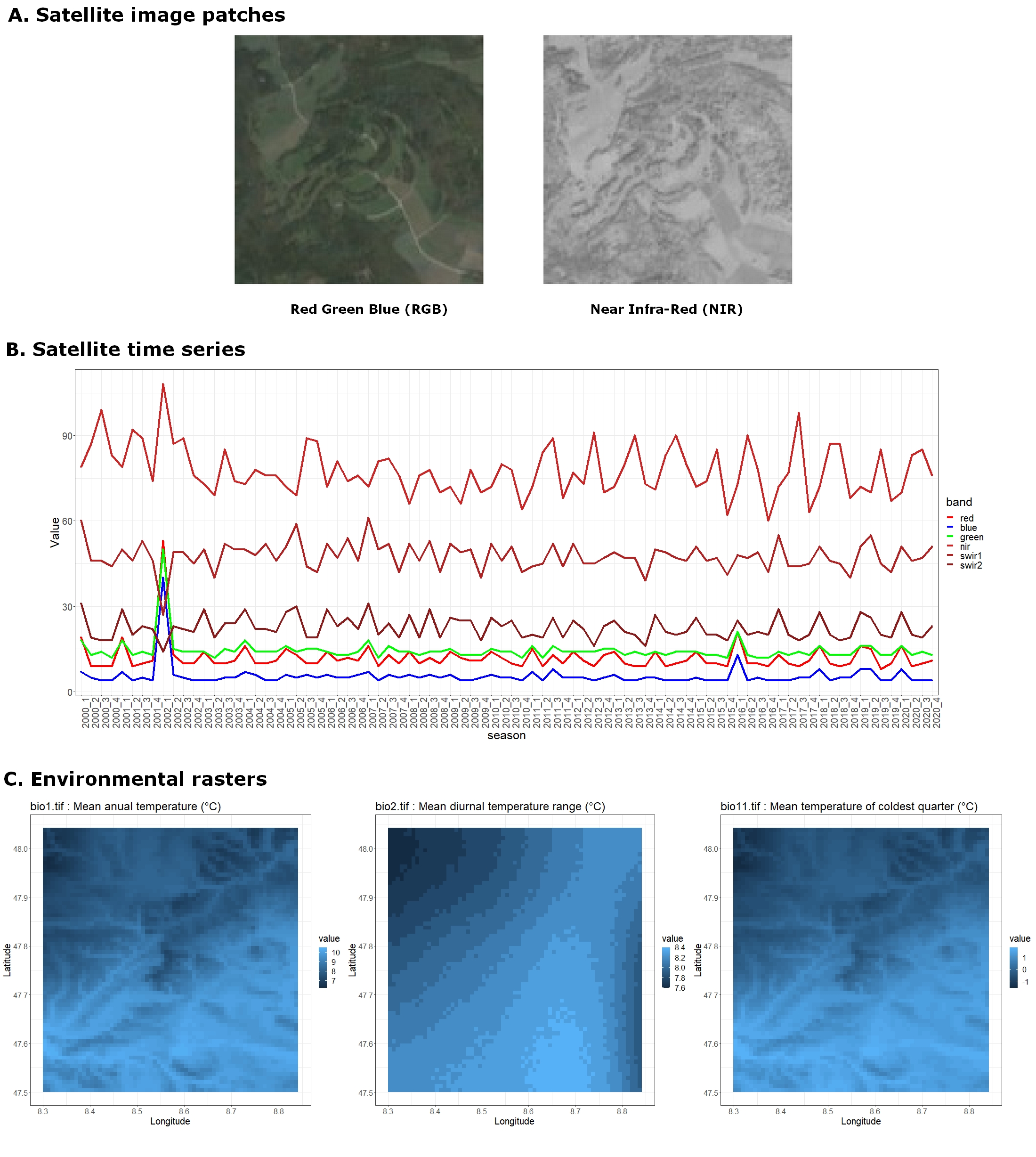
- Satellite image patches: 3-band (RGB) and 1-band (NIR) 128x128 images at 10m resolution.
- Satellite time series: Up to 20 years of values for six satellite bands (R, G, B, NIR, SWIR1, and SWIR2).
- Environmental rasters Various climatic, pedologic, land use, and human footprint variables at the European scale. We provide scalar values, time-series, and original rasters from which you may extract local 2D images.
There are three separate folders with the relevant data on the Seafile available for training. The detailed description is provided below and again on SeaFile in separate "Readme" files in relevant folders.
Links
❗New Seafile repository❗: repository containing all the data. To optimize download times, see the section data downloading at the bottom of this page.
❗GLC GitHub repository❗: Useful codes to manipulate data with simple data loaders, examples, and sample data. More dataloaders can be added after the challenge starts.
CVPR24 and CLEF24 Context
This competition is held jointly as part of:
- the LifeCLEF 2024 lab of the CLEF 2024 conference, and of
- the FGVC11 workshop organized in conjunction with CVPR 2024 conference.
Being part of scientific research, the participants are encouraged to participate to both events.
In particular, only participants who submitted a working note paper to LifeCLEF (see below) will be part of the officially published ranking used for scientific communication.
CEUR Working Notes submission
For detailed instructions, please refer to SUBMISSION INSTRUCTIONS.
A summary of the most important points:
- All participating teams with at least one graded submission, regardless of the score, should submit a CEUR working notes paper.
- Submission of reports is done through EasyChair – please make absolutely sure that the author (names and order), title, and affiliation information you provide in EasyChair match the submitted PDF exactly
- Deadline for the submission of initial CEUR-WS Working Notes Papers (for the peer-review process): 31 May 2024
- Deadline for the submission of Camera Ready CEUR-WS Working Notes Papers:: 8 July 2024
- Templates are available here
- Working Notes Papers should cite both the LifeCLEF 2024 overview paper as well as the PlantCLEF task overview paper, citation information will be added in the Citations section below as soon as the titles have been finalized.
Credit
This project has received funding from the European Union’s Horizon research and innovation program under grant agreement No 101060639 (MAMBO project) and No 101060693 (GUARDEN project).


