- ImageCLEF 2026
- LifeCLEF 2026
- ImageCLEF 2025
- LifeCLEF 2025
- ImageCLEF 2024
- LifeCLEF 2024
- ImageCLEF 2023
- LifeCLEF 2023
- ImageCLEF 2022
- LifeCLEF2022
- ImageCLEF 2021
- LifeCLEF 2021
- ImageCLEF 2020
- LifeCLEF 2020
- ImageCLEF 2019
- LifeCLEF 2019
- ImageCLEF 2018
- LifeCLEF 2018
- ImageCLEF 2017
- LifeCLEF2017
- ImageCLEF 2016
- LifeCLEF 2016
- ImageCLEF 2015
- LifeCLEF 2015
- ImageCLEF 2014
- LifeCLEF 2014
- ImageCLEF 2013
- ImageCLEF 2012
- ImageCLEF 2011
- ImageCLEF 2010
- ImageCLEF 2009
- ImageCLEF 2008
- ImageCLEF 2007
- ImageCLEF 2006
- ImageCLEF 2005
- ImageCLEF 2004
- ImageCLEF 2003
- Publications
- Old resources
You are here
ImageCLEFmedical Caption
Welcome to the 7th edition of the Caption Task!
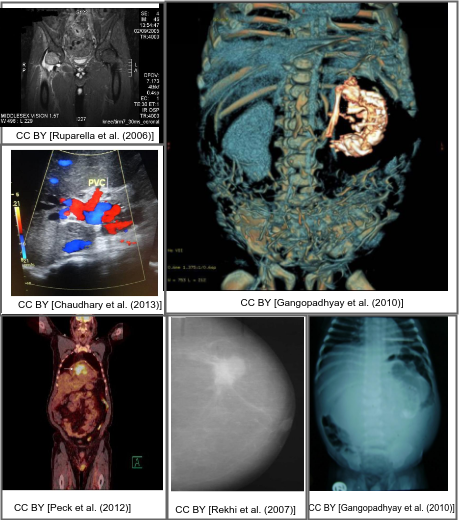
Motivation
Interpreting and summarizing the insights gained from medical images such as radiology output is a time-consuming task that involves highly trained experts and often represents a bottleneck in clinical diagnosis pipelines.
Consequently, there is a considerable need for automatic methods that can approximate this mapping from visual information to condensed textual descriptions. The more image characteristics are known, the more structured are the radiology scans and hence, the more efficient are the radiologists regarding interpretation. We work on the basis of a large-scale collection of figures from open access biomedical journal articles (PubMed Central). All images in the training data are accompanied by UMLS concepts extracted from the original image caption.
Lessons learned:
- In the first and second editions of this task, held at ImageCLEF 2017 and ImageCLEF 2018, participants noted a broad variety of content and situation among training images. In 2019, the training data was reduced solely to radiology images, with ImageCLEF 2020 adding additional imaging modality information, for pre-processing purposes and multi-modal approaches.
- The focus in ImageCLEF 2021 lay in using real radiology images annotated by medical doctors. This step aimed at increasing the medical context relevance of the UMLS concepts, but more images of such high quality are difficult to acquire.
- As uncertainty regarding additional source was noted, we will clearly separate systems using exclusively the official training data from those that incorporate additional sources of evidence
- For ImageCLEF 2022, an extended version of the ImageCLEF 2020 dataset was used. There were several issues with the dataset (large number of concepts, lemmatization errors, duplicate captions), which will be tackled for the 7th edition of the task, alongside an updated primary evaluation metric for the caption prediction subtask.
News
- 12.10.2022: website goes live
- 12.01.2023: registration opens
- 09.02.2023: development dataset released
- 14.03.2023: test dataset released
- 09.05.2023: run submission deadline extended to May 12
- 19.05.2023: results published
Task Description
For captioning, participants will be requested to develop solutions for automatically identifying individual components from which captions are composed in Radiology Objects in COntext images.
ImageCLEFmedical Caption 2023 consists of two substaks:
- Concept Detection Task
- Caption Prediction Task
Concept Detection Task
The first step to automatic image captioning and scene understanding is identifying the presence and location of relevant concepts in a large corpus of medical images. Based on the visual image content, this subtask provides the building blocks for the scene understanding step by identifying the individual components from which captions are composed. The concepts can be further applied for context-based image and information retrieval purposes.
Evaluation is conducted in terms of set coverage metrics such as precision, recall, and combinations thereof.
Caption Prediction Task
On the basis of the concept vocabulary detected in the first subtask as well as the visual information of their interaction in the image, participating systems are tasked with composing coherent captions for the entirety of an image. In this step, rather than the mere coverage of visual concepts, detecting the interplay of visible elements is crucial for strong performance.
This year, we will use BERTScore as the primary evaluation metric and ROUGE as the secondary evaluation metric for the caption prediction subtask. Other metrics such as METEOR, CIDEr, and BLEU will also be published.
Data
The data for the caption task will contain curated images from the medical literature including their captions and associated UMLS terms that are manually controlled as metadata. A more diverse data set will be made available to foster more complex approaches.
An updated and extended version of the Radiology Objects in COntext (ROCO) dataset [1] is used for both subtasks. As in previous editions, the dataset originates from biomedical articles of the PMC OpenAccess subset.
Training Set: Consists of 60,918 radiology images
Validation Set: Consists of 10,437 radiology images
Test Set: Consists of 10,473 radiology images
Concept Detection Task
The concepts were generated using a reduced subset of the UMLS 2022 AB release. To improve the feasibility of recognizing concepts from the images, concepts were filtered based on their semantic type. Concepts with low frequency were also removed, based on suggestions from previous years.
Caption Prediction Task
For this task each caption is pre-processed in the following way:
- removal of links from the captions
Evaluation methodology
For assessing performance, classic metrics will be used, ranging from F1 score, accuracy, and BERTScore or other measures like ROUGE to measure text similarity.
The source code of the evaluation script is available at GitHub (private repository only available to participants).
Concept Detection
Evaluation is conducted in terms of F1 scores between system predicted and ground truth concepts, using the following methodology and parameters:
- The default implementation of the Python scikit-learn (v0.17.1-2) F1 scoring method is used. It is documented here.
- A Python (3.x) script loads the candidate run file, as well as the ground truth (GT) file, and processes each candidate-GT concept sets
- For each candidate-GT concept set, the y_pred and y_true arrays are generated. They are binary arrays indicating for each concept contained in both candidate and GT set if it is present (1) or not (0).
- The F1 score is then calculated. The default 'binary' averaging method is used.
- All F1 scores are summed and averaged over the number of elements in the test set (7'601), giving the final score.
The ground truth for the test set was generated based on the same reduced subset of the UMLS 2022 AB release which was used for the training data (see above for more details).
Caption Prediction
This year, BERTScore is used as the primary metric instead of BLEU, and ROUGE is used as a secondary metric. Other metrics like METEOR and CIDR will be reported after the challenge concludes.
For all metrics, each caption is pre-processed in the same way:
- The caption is converted to lower-case
- Replace numbers with the token 'number'
- Remove punctuation
BERTScore is calculated using the following methodology and parameters:
The native Python implementation of BERTScore is used, which can be found in the GitHub repository here. This scoring method is based on the paper "BERTScore: Evaluating Text Generation with BERT" and aims to measure the quality of generated text by comparing it to a reference.
To calculate BERTScore, we use the microsoft/deberta-xlarge-mnli model, which can be found on the Hugging Face Model Hub. The model is pretrained on a large corpus of text and fine-tuned for natural language inference tasks. It can be used to compute contextualized word embeddings, which are essential for BERTScore calculation.
To compute the final BERTScore, we first calculate the individual score (F1) for each sentence. Then all BERTScores are summed and averaged over the number of captions, yielding the final score.
The ROUGE scores are calculated using the following methodology and parameters:
- The native python implementation of ROUGE scoring method is used. It is designed to replicate results from the original perl package that was introduced in the original article describing the ROUGE evaluation method.
- Specifically, we calculate the ROUGE-1 (F-measure) score, which measures the number of matching unigrams between the model-generated text and a reference.
- A Python (3.7) script loads the candidate run file, as well as the ground truth (GT) file, and processes each candidate-GT caption pair
- The ROUGE score is then calculated. Note that the caption is always considered as a single sentence, even if it actually contains several sentences.
- All ROUGE scores are summed and averaged over the number of captions, giving the final score.
Additional metrics were mainly calculated using the pycocoevalcap [pypi.org] library.
- For the METEOR score, the v1.5 jar is used for the calculation, which was introduced in Meteor Universal: Language Specific Translation Evaluation for Any Target Language [aclanthology.org]
- CIDEr score is based on CIDEr: Consensus-Based Image Description Evaluation [cv-foundation.org]
- The java implementation of SPICE introduced in SPICE.pdf [panderson.me] is used.
- For BLEU, BLEU-1 from pycocoevalcap is used
- BLEURT: A BERT-based metric for evaluating text generation, using a model specifically trained to align with human judgements. [2004.04696] BLEURT: Learning Robust Metrics for Text Generation (arxiv.org). GitHub: google-research/bleurt: BLEURT is a metric for Natural Language Generation based on transfer learning. (github.com) with BLEURT-20 Model
- CLIPScore: An evaluation metric for multimodal models that measures the coherence between an image and a caption, based on OpenAI's CLIP model. [2104.08718] CLIPScore: A Reference-free Evaluation Metric for Image Captioning (arxiv.org). GitHub: jmhessel/clipscore: CLIPScore EMNLP code (github.com)
Participant registration
Please refer to the general ImageCLEF registration instructions
Preliminary Schedule
- 19 December 2022: Registration opens
- 6 Feburary 2023: Release of the training and validation sets
- 14 March 2023: Release of the test sets
- 22 April 2023: Registration closes
-
10 May12 May 2023: Run submission deadline -
17 May19 May 2023: Release of the processed results by the task organizers - 5 June 2023: Submission of participant papers [CEUR-WS]
- 23 June 2023: Notification of acceptance
- 7 July 2023: Camera ready copy of participant papers and extended lab overviews [CEUR-WS]
- 18-21 September 2023: CLEF 2023, Thessaloniki, Greece
Submission Instructions
Please find detailed submission instructions on GitHub (private repository accessible only for participants).
Results
The tables below contain only the best runs of each team, for a complete list of all runs please see the CSV files in this Sciebo folder.
Concept Detection Task
For the cconcept detection task, the ranking is based on the F1-score as described in the Evaluation Methodologies section above. Additionally, a F1-score Manual was calculated using a subset of manually validated concepts (anatomy, directionality, and image modality) only.
| Team Name | Run ID | F1-Score | F1-Score Manual |
|---|---|---|---|
| AUEB-NLP-Group | 4 | 0.522272 | 0.925842 |
| KDE-Lab_Med | 10 | 0.507414 | 0.932091 |
| VCMI | 8 | 0.499812 | 0.916184 |
| IUST_NLPLAB | 7 | 0.495863 | 0.880381 |
| Clef-CSE-GAN-Team | 1 | 0.495730 | 0.910585 |
| CS_Morgan | 2 | 0.483401 | 0.890151 |
| SSNSheerinKavitha | 1 | 0.464894 | 0.860296 |
| closeAI2023 | 5 | 0.448105 | 0.856928 |
| SSN_MLRG | 3 | 0.017250 | 0.112211 |
Caption Prediction Task
For the caption prediction task, the ranking is based on the BERTScore.
| Team Name | Run ID | BERTScore | ROUGE | BLEURT | BLEU | METEOR | CIDEr | CLIPScore |
|---|---|---|---|---|---|---|---|---|
| CSIRO | 4 | 0.642519 | 0.244618 | 0.313707 | 0.161486 | 0.079775 | 0.202512 | 0.814717 |
| closeAI2023 | 7 | 0.628106 | 0.240061 | 0.320915 | 0.184624 | 0.087254 | 0.237704 | 0.807454 |
| AUEB-NLP-Group | 2 | 0.617034 | 0.213014 | 0.295011 | 0.169212 | 0.071982 | 0.146601 | 0.803888 |
| PCLmed | 5 | 0.615190 | 0.252756 | 0.316561 | 0.217150 | 0.092063 | 0.231535 | 0.802123 |
| VCMI | 5 | 0.614736 | 0.217545 | 0.308386 | 0.165322 | 0.073449 | 0.172042 | 0.808184 |
| KDE-Lab_Med | 3 | 0.614538 | 0.222341 | 0.301391 | 0.156465 | 0.072441 | 0.181853 | 0.806207 |
| SSN_MLRG | 1 | 0.601933 | 0.211177 | 0.277434 | 0.141797 | 0.061514 | 0.128443 | 0.775915 |
| DLNU_CCSE | 1 | 0.600546 | 0.202888 | 0.262998 | 0.105948 | 0.055716 | 0.133207 | 0.772518 |
| CS_Morgan | 10 | 0.581949 | 0.156419 | 0.224238 | 0.056632 | 0.043649 | 0.083982 | 0.759258 |
| Clef-CSE-GAN-Team | 2 | 0.581625 | 0.218103 | 0.269043 | 0.145035 | 0.070155 | 0.173664 | 0.789327 |
| Bluefield-2023 | 3 | 0.577966 | 0.153448 | 0.271642 | 0.154316 | 0.060069 | 0.100910 | 0.783725 |
| IUST_NLPLAB | 6 | 0.566886 | 0.289774 | 0.222957 | 0.268452 | 0.100354 | 0.177266 | 0.806763 |
| SSNSheerinKavitha | 4 | 0.544106 | 0.086648 | 0.215170 | 0.074905 | 0.025768 | 0.014313 | 0.687312 |
CEUR Working Notes
For detailed instructions, please refer to this PDF file. A summary of the most important points:
- All participating teams with at least one graded submission, regardless of the score, should submit a CEUR working notes paper.
- Teams who participated in both tasks should generally submit only one report
- Submission of reports is done through EasyChair – please make absolutely sure that the author (names and order), title, and affiliation information you provide in EasyChair match the submitted PDF exactly!
- Strict deadline for Working Notes Papers: 05 June 2023 (23:59 CEST)
- Strict deadline for CEUR-WS Camera Ready Working Notes Papers: 07 July 2023 (23:59 CEST)
- Make sure to include the signed Copyright Form
- Templates are available here
- Working Notes Papers should cite both the ImageCLEF 2023 overview paper as well as the ImageCLEFmedical task overview paper, citation information is available in the Citations section below.
Citations
When referring to ImageCLEF 2023, please cite the following publication:
- Bogdan Ionescu, Henning Müller, Ana-Maria Drăgulinescu, Wen-wai Yim, Asma Ben Abacha, Neal Snider, Griffin Adams, Meliha Yetisgen, Johannes Rückert, Alba García Seco de Herrera, Christoph M. Friedrich, Louise Bloch, Raphael Brüngel, Ahmad Idrissi-Yaghir, Henning Schäfer, Steven A. Hicks, Michael A. Riegler, Vajira Thambawita, Andrea Storås, Pål Halvorsen, Nikolaos Papachrysos, Johanna Schöler, Debesh Jha, Alexandra-Georgiana Andrei, Ahmedkhan Radzhabov, Ioan Coman, Vassili Kovalev, Alexandru Stan, George Ioannidis, Hugo Manguinhas, Liviu-Daniel Ștefan, Mihai Gabriel Constantin, Mihai Dogariu, Jérôme Deshayes, Adrian Popescu, Overview of the ImageCLEF 2023: Multimedia Retrieval in Medical, Social Media and Recommender Systems Applications, in Experimental IR Meets Multilinguality, Multimodality, and Interaction.Proceedings of the 14th International Conference of the CLEF Association (CLEF 2023), Springer Lecture Notes in Computer Science LNCS, Thessaloniki, Greece, September 18-21, 2023.
- BibTex:
@inproceedings{ImageCLEF2023,
author = {Bogdan Ionescu and Henning M\"uller and Ana{-}Maria Dr\u{a}gulinescu and Wen{-}wai Yim and Asma {Ben Abacha} and Neal Snider and Griffin Adams and Meliha Yetisgen and Johannes R\"uckert and Alba {Garc\’{\i}a Seco de Herrera} and Christoph M. Friedrich and Louise Bloch and Raphael Br\"ungel and Ahmad Idrissi{-}Yaghir and Henning Sch\"afer and Steven A. Hicks and Michael A. Riegler and Vajira Thambawita and Andrea Stor{\r a}s and P{\r a}l Halvorsen and Nikolaos Papachrysos and Johanna Sch\"oler and Debesh Jha and Alexandra{-}Georgiana Andrei and Ahmedkhan Radzhabov and Ioan Coman and Vassili Kovalev and Alexandru Stan and George Ioannidis and Hugo Manguinhas and Liviu{-}Daniel \c{S}tefan and Mihai Gabriel Constantin and Mihai Dogariu and J\'er\^ome Deshayes and Adrian Popescu},
title = {{Overview of ImageCLEF 2023}: Multimedia Retrieval in Medical, SocialMedia and Recommender Systems Applications},
booktitle = {Experimental IR Meets Multilinguality, Multimodality, and Interaction},
series = {Proceedings of the 14th International Conference of the CLEF Association (CLEF 2023)},
year = {2023},
publisher = {Springer Lecture Notes in Computer Science LNCS},
pages = {},
month = {September 18-21},
address = {Thessaloniki, Greece}
}
When referring to ImageCLEFmedical 2023 Caption general goals, general results, etc. please cite the following publication:
- Johannes Rückert, Asma Ben Abacha, Alba G. Seco de Herrera, Louise Bloch, Raphael Brüngel, Ahmad Idrissi-Yaghir, Henning Schäfer, Henning Müller and Christoph M. Friedrich. Overview of ImageCLEFmedical 2023 – Caption Prediction and Concept Detection, in Experimental IR Meets Multilinguality, Multimodality, and Interaction. CEUR Workshop Proceedings (CEUR-WS.org), Thessaloniki, Greece, September 18-21, 2023.
- BibTex:
@inproceedings{ImageCLEFmedicalCaptionOverview2023,
author = {R\"uckert, Johannes and Ben Abacha, Asma and G. Seco de Herrera, Alba and Bloch, Louise and Br\"ungel, Raphael and Idrissi-Yaghir, Ahmad and Sch\"afer, Henning and M\"uller, Henning and Friedrich, Christoph M.},
title = {Overview of {ImageCLEFmedical} 2023 -- {Caption Prediction and Concept Detection}},
booktitle = {CLEF2023 Working Notes},
series = {{CEUR} Workshop Proceedings},
year = {2023},
volume = {},
publisher = {CEUR-WS.org},
pages = {},
month = {September 18-21},
address = {Thessaloniki, Greece}
}
Contact
Organizers:
- Johannes Rückert <johannes.rueckert(at)fh-dortmund.de>, University of Applied Sciences and Arts Dortmund, Germany
- Asma Ben Abacha <abenabacha(at)microsoft.com>, Microsoft, USA
- Alba García Seco de Herrera <alba.garcia(at)essex.ac.uk>,University of Essex, UK
- Christoph M. Friedrich <christoph.friedrich(at)fh-dortmund.de>, University of Applied Sciences and Arts Dortmund, Germany
- Henning Müller <henning.mueller(at)hevs.ch>, University of Applied Sciences Western Switzerland, Sierre, Switzerland
- Louise Bloch <louise.bloch(at)fh-dortmund.de>, University of Applied Sciences and Arts Dortmund, Germany
- Raphael Brüngel <raphael.bruengel(at)fh-dortmund.de>, University of Applied Sciences and Arts Dortmund, Germany
- Ahmad Idrissi-Yaghir <ahmad.idrissi-yaghir(a)fh-dortmund.de>, University of Applied Sciences and Arts Dortmund, Germany
- Henning Schäfer <henning.schaefer(at)uk-essen.de>, University of Applied Sciences and Arts Dortmund, Germany
Acknowledgments
[1] O. Pelka, S. Koitka, J. Rückert, F. Nensa und C. M. Friedrich „Radiology Objects in COntext (ROCO): A Multimodal Image Dataset“, Proceedings of the MICCAI Workshop on Large-scale Annotation of Biomedical data and Expert Label Synthesis (MICCAI LABELS 2018), Granada, Spain, September 16, 2018, Lecture Notes in Computer Science (LNCS) Volume 11043, Page 180-189, DOI: 10.1007/978-3-030-01364-6_20, Springer Verlag, 2018.

